Island Jacobian Analysis: Periodic Orbits and Monodromy Matrices¶
This notebook demonstrates the full Jacobian/monodromy analysis for magnetic island chains in an analytic stellarator equilibrium, ported from the Julia MCF_scripts approach.
Scientific background¶
A periodic field-line orbit (cycle) is a field line that returns to its starting point after exactly \(n\) toroidal turns: \(X(\phi + 2\pi n) = X(\phi)\).
These are fixed points of the \(n\)-th iterate of the Poincaré map \(P^n\). For a resonance \(q = m/n\) (with \(q = B_\phi r / (B_{pol} R)\), the orbit period is \(m\) turns.
The Jacobian matrix \(DX(\phi)\) evolves as:
where \(A_{ij} = \partial(R B_{pol,i}/B_\phi) / \partial x_j\).
The monodromy matrix \(M = DP(\phi_0 + 2\pi n)\) gives the linearized \(n\)-turn map. Eigenvalues of \(M\):
\(|\lambda| = 1\): elliptic (O-point, stable)
\(|\lambda| > 1\): hyperbolic (X-point, unstable)
[1]:
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.insert(0, '../..')
from pyna.toroidal.equilibrium.stellarator import SimpleStellarartor
from pyna.topo.toroidal_cycle import (
poincare_map_n, poincare_map_n_trajectory,
jacobian_of_poincare_map, find_cycle, find_all_cycles_near_resonance,
ToroidalPeriodicOrbitTrace as PeriodicOrbit,
)
from pyna.topo.monodromy import (
compute_Jac, build_A_matrix_func, build_delta_b_pol_func,
orbit_shift_under_perturbation, monodromy_change_under_perturbation,
MonodromyAnalysis,
)
plt.rcParams['figure.figsize'] = (10, 7)
plt.rcParams['font.size'] = 12
print('Imports OK')
Imports OK
1. Setup: Build SimpleStellarartor with q=2/1 island chain¶
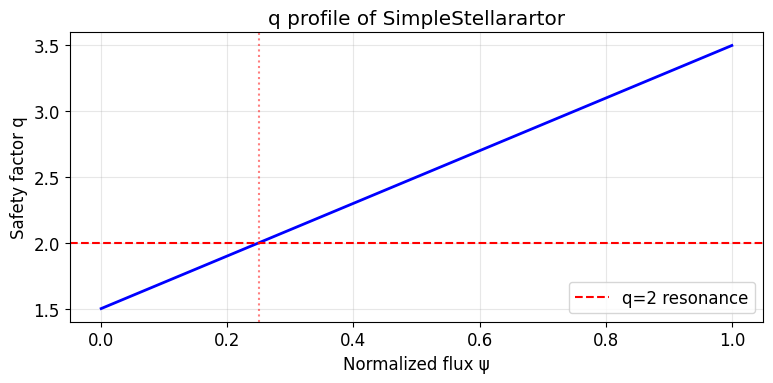
We use m_h=2, n_h=1, epsilon_h=0.02 to create a q=2 resonance. In this model, q = m_h/n_h = 2, and the island chain has period 2 (2 toroidal turns).
The q profile: q(r) = q0 + (q1-q0)*psi = q0 + (q1-q0)*(r/r0)²
At r=0: q = q0 = 1.5
At r=r0: q = q1 = 3.5
q=2 surface at: psi = (q-q0)/(q1-q0) = 0.5/2.0 = 0.25 ?r/r0 = 0.5
[2]:
# Build the stellarator
epsilon_h = 0.02 # helical ripple amplitude
stellarator = SimpleStellarartor(
R0=3.0, r0=0.35, B0=1.0,
q0=1.5, q1=3.5,
m_h=2, n_h=1, epsilon_h=epsilon_h,
)
R0, r0 = stellarator.R0, stellarator.r0
field_func = stellarator.field_func
# Domain limits
RZlimit = (R0 - r0*1.5, R0 + r0*1.5, -r0*1.5, r0*1.5)
# Find q=2/1 resonant surface
psi_list = stellarator.resonant_psi(2, 1)
psi_res = psi_list[0]
r_res = np.sqrt(psi_res) * r0
print(f"q=2/1 resonance at psi={psi_res:.3f}, r_res={r_res:.4f} m")
# q profile plot
psi_arr = np.linspace(0, 1, 100)
q_arr = [stellarator.q_of_psi(p) for p in psi_arr]
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
ax.plot(psi_arr, q_arr, 'b-', linewidth=2)
ax.axhline(2.0, color='r', linestyle='--', label='q=2 resonance')
ax.axvline(psi_res, color='r', linestyle=':', alpha=0.5)
ax.set_xlabel('Normalized flux ψ')
ax.set_ylabel('Safety factor q')
ax.set_title('q profile of SimpleStellarartor')
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('q_profile.png', dpi=100, bbox_inches='tight')
plt.show()
print('q profile plot saved')
q=2/1 resonance at psi=0.250, r_res=0.1750 m
q profile plot saved
2. Poincaré Section and Island Chain Visualization¶
[3]:
# Compute Poincaré section
print('Computing Poincaré section (this may take ~30s)...')
# Trace field lines from various starting points
n_field_lines = 8
n_poincare_turns = 100
poincare_points = [] # (R, Z) lists per field line
for i, r_frac in enumerate(np.linspace(0.15, 0.95, n_field_lines)):
r_start = r_frac * r0
R_start = R0 + r_start
Z_start = 0.0
Rs, Zs = [R_start], [Z_start]
R_curr, Z_curr = R_start, Z_start
for _ in range(n_poincare_turns):
R_next, Z_next = poincare_map_n(
field_func, [R_curr, Z_curr, 0.0], n_turns=1, dt=0.15, RZlimit=RZlimit
)
if np.isnan(R_next):
break
Rs.append(R_next)
Zs.append(Z_next)
R_curr, Z_curr = R_next, Z_next
poincare_points.append((np.array(Rs), np.array(Zs)))
print(f' Field line {i+1}/{n_field_lines}: {len(Rs)} points')
print('Poincaré section computed')
Computing Poincaré section (this may take ~30s)...
Field line 1/8: 101 points
Field line 2/8: 101 points
Field line 3/8: 101 points
Field line 4/8: 84 points
Field line 5/8: 3 points
Field line 6/8: 2 points
Field line 7/8: 2 points
Field line 8/8: 2 points
Poincaré section computed
[4]:
# Find O-points and X-points using Newton-Raphson
print('Finding X/O points...')
# Known approximate locations from scan (for epsilon_h=0.02, q=2/1)
# X-points: near theta=±0.79 and ±2.36 at r=r_res
# O-points: near theta=±2.32 at r=r_res (for small epsilon)
cycle_seeds = [
# X-point candidates
np.array([R0 + r_res*np.cos(0.79), r_res*np.sin(0.79), 0.0]),
np.array([R0 + r_res*np.cos(-0.79), r_res*np.sin(-0.79), 0.0]),
np.array([R0 + r_res*np.cos(2.36), r_res*np.sin(2.36), 0.0]),
np.array([R0 + r_res*np.cos(-2.36), r_res*np.sin(-2.36), 0.0]),
# O-point candidates (scan found dist? at theta?.02, r=0.1625)
np.array([R0 + r_res*np.cos(3.02), r_res*np.sin(3.02), 0.0]),
np.array([R0 + r_res*np.cos(-2.32), r_res*np.sin(-2.32), 0.0]),
np.array([R0 + r_res*np.cos(2.32), r_res*np.sin(2.32), 0.0]),
np.array([R0 + r_res*np.cos(0.25), r_res*np.sin(0.25), 0.0]),
]
found_orbits = []
for seed in cycle_seeds:
orbit = find_cycle(
field_func, seed, n_turns=2, dt=0.15, RZlimit=RZlimit,
max_iter=30, tol=1e-8, n_fallback_seeds=4, fallback_radius=0.02,
)
if orbit is None:
continue
dist_axis = np.sqrt((orbit.rzphi0[0]-R0)**2 + orbit.rzphi0[1]**2)
if dist_axis < 0.05:
continue # skip axis fixed point
# Deduplicate
dup = any(np.linalg.norm(orbit.rzphi0[:2]-fo.rzphi0[:2]) < 1e-4 for fo in found_orbits)
if not dup:
found_orbits.append(orbit)
o_points = [o for o in found_orbits if o.is_stable]
x_points = [o for o in found_orbits if not o.is_stable]
print(f'Found {len(o_points)} O-points and {len(x_points)} X-points')
for o in o_points:
print(f' O-point: ({o.rzphi0[0]:.4f}, {o.rzphi0[1]:.4f}), k={o.stability_index:.4f}')
for x in x_points:
print(f' X-point: ({x.rzphi0[0]:.4f}, {x.rzphi0[1]:.4f}), k={x.stability_index:.4f}')
Finding X/O points...
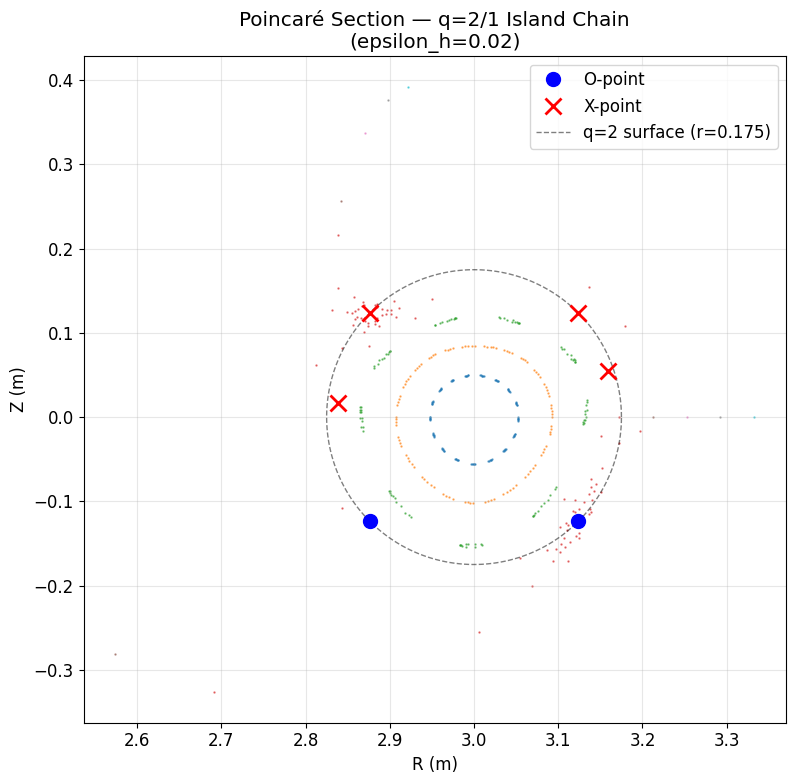
Found 2 O-points and 4 X-points
O-point: (3.1237, -0.1237), k=-0.1045
O-point: (2.8763, -0.1237), k=0.3026
X-point: (3.1237, 0.1237), k=0.3024
X-point: (2.8763, 0.1237), k=-0.1042
X-point: (2.8383, 0.0169), k=2.0324
X-point: (3.1587, 0.0546), k=2.0320
[5]:
# Plot Poincaré section with X/O points marked
fig, ax = plt.subplots(figsize=(10, 8))
colors = plt.cm.tab10(np.linspace(0, 0.9, n_field_lines))
for i, (Rs, Zs) in enumerate(poincare_points):
ax.scatter(Rs, Zs, s=0.5, color=colors[i], alpha=0.6)
# Mark O-points (blue circles)
for o in o_points:
ax.plot(o.rzphi0[0], o.rzphi0[1], 'bo', markersize=10,
label='O-point' if o is o_points[0] else '', zorder=5)
# Mark X-points (red x's)
for x in x_points:
ax.plot(x.rzphi0[0], x.rzphi0[1], 'rx', markersize=12, markeredgewidth=2,
label='X-point' if x is x_points[0] else '', zorder=5)
# Mark resonant surface
theta_arr = np.linspace(0, 2*np.pi, 200)
ax.plot(R0 + r_res*np.cos(theta_arr), r_res*np.sin(theta_arr), 'k--',
linewidth=1, alpha=0.5, label=f'q=2 surface (r={r_res:.3f})')
ax.set_xlabel('R (m)')
ax.set_ylabel('Z (m)')
ax.set_title(f'Poincaré Section ?q=2/1 Island Chain\n(epsilon_h={epsilon_h})')
ax.set_aspect('equal')
ax.legend(loc='upper right')
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('poincare_section.png', dpi=100, bbox_inches='tight')
plt.show()
print('Poincaré section saved')
# Note: For interactive Plotly version, use:
# import plotly.graph_objects as go
# fig = go.Figure()
# for Rs, Zs in poincare_points:
# fig.add_trace(go.Scatter(x=Rs, y=Zs, mode='markers', marker=dict(size=1)))
# for o in o_points:
# fig.add_trace(go.Scatter(x=[o.rzphi0[0]], y=[o.rzphi0[1]], mode='markers',
# marker=dict(size=10, symbol='circle', color='blue')))
# fig.show()
Poincaré section saved
3. Monodromy Analysis for O-point¶
The monodromy matrix \(M = DP(\phi_0 + 2\pi n)\) encodes the stability of the periodic orbit. For an O-point (elliptic), eigenvalues lie on the unit circle: \(|\lambda| = 1\).
Greene’s residue: \(R_G = (2 - \text{Tr}(M))/4\)
\(0 < R_G < 1\): elliptic (stable)
\(R_G < 0\): hyperbolic (unstable, standard)
\(R_G > 1\): hyperbolic with reflection
[6]:
# Pick the first O-point for analysis
if not o_points:
print('No O-points found ?using first found orbit for demo')
opoint = found_orbits[0] if found_orbits else None
else:
opoint = o_points[0]
if opoint is None:
print('No orbits found to analyze')
else:
print(f'Analyzing O-point at ({opoint.rzphi0[0]:.4f}, {opoint.rzphi0[1]:.4f})')
print(f' stability_index = {opoint.stability_index:.6f}')
print(f' eigenvalues = {opoint.eigenvalues}')
# Compute full monodromy analysis
print('Computing monodromy (variational equations)...')
monodromy_O = compute_Jac(
field_func, opoint, dt_output=0.1, rtol=1e-8, atol=1e-9
)
print(f' Monodromy matrix M:')
print(f' {monodromy_O.Jac}')
print(f' det(M) = {np.linalg.det(monodromy_O.Jac):.8f} (should be ?1)')
print(f' Tr(M)/2 = {monodromy_O.stability_index:.6f}')
print(f" Greene's residue = {monodromy_O.Greene_residue:.6f}")
print(f' Eigenvalues = {monodromy_O.eigenvalues}')
Analyzing O-point at (3.1237, -0.1237)
stability_index = -0.104476
eigenvalues = [-0.10447603+0.99401493j -0.10447603-0.99401493j]
Computing monodromy (variational equations)...
Monodromy matrix M:
[[-1.95431995 1.21064224]
[-3.64518199 1.74639332]]
det(M) = 0.99999999 (should be ?1)
Tr(M)/2 = -0.103963
Greene's residue = 0.551982
Eigenvalues = [-0.10396332+0.99458113j -0.10396332-0.99458113j]
[7]:
if opoint is not None and 'monodromy_O' in dir():
phi_arr = monodromy_O.phi_arr
# Compute eigenvalues and det along orbit
DP_eigvals = np.array([np.linalg.eigvals(DP) for DP in monodromy_O.J_arr])
DP_dets = np.array([np.linalg.det(DP) for DP in monodromy_O.J_arr])
fig, axes = plt.subplots(3, 1, figsize=(10, 10))
# Plot |eigenvalues| of J along orbit
ax = axes[0]
ax.plot(phi_arr/(2*np.pi), np.abs(DP_eigvals[:, 0]), 'b-', label='|λ?φ)|')
ax.plot(phi_arr/(2*np.pi), np.abs(DP_eigvals[:, 1]), 'r--', label='|λ?φ)|')
ax.axhline(1.0, color='k', linestyle=':', alpha=0.5)
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('|eigenvalue of DP|')
ax.set_title('O-point: Eigenvalue evolution along orbit')
ax.legend()
ax.grid(True, alpha=0.3)
# Plot det(J) ?should stay ?1
ax = axes[1]
ax.plot(phi_arr/(2*np.pi), DP_dets, 'g-', linewidth=2)
ax.axhline(1.0, color='k', linestyle=':', alpha=0.5, label='det=1 (area-preserving)')
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('det(J(φ))')
ax.set_title('Area preservation check: det(J) ?1')
ax.legend()
ax.grid(True, alpha=0.3)
# Plot Greene residue evolution (Tr(J)/2)
ax = axes[2]
Tr_half = np.array([np.trace(J)/2 for J in monodromy_O.J_arr])
ax.plot(phi_arr/(2*np.pi), Tr_half, 'm-', linewidth=2)
ax.axhline(0.0, color='k', linestyle=':', alpha=0.3)
ax.axhline(monodromy_O.stability_index, color='r', linestyle='--',
label=f'Final Tr(M)/2 = {monodromy_O.stability_index:.4f}')
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('Tr(J(φ))/2')
ax.set_title('Stability index evolution along O-point orbit')
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('monodromy_opoint.png', dpi=100, bbox_inches='tight')
plt.show()
print('Monodromy O-point plot saved')
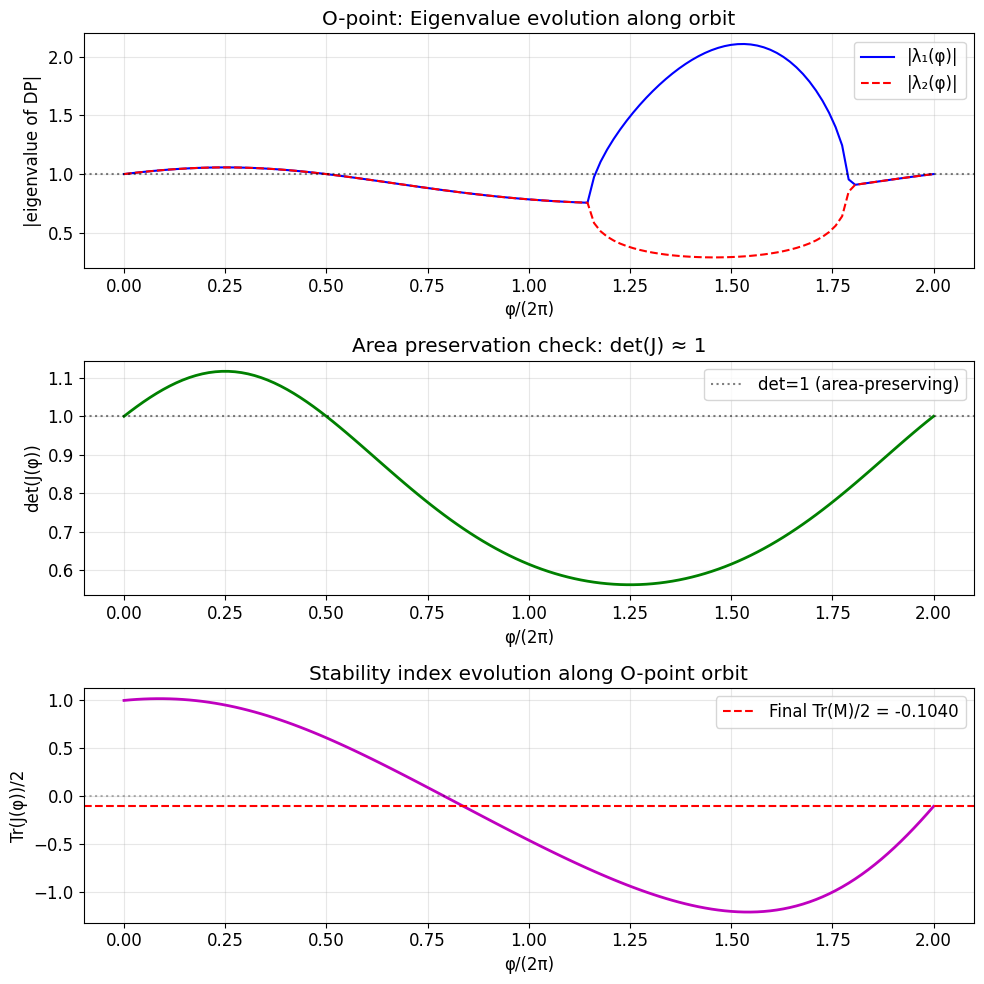
Monodromy O-point plot saved
4. Monodromy Analysis for X-point¶
For an X-point (hyperbolic), eigenvalues are real and \(|\lambda| > 1\). Nearby orbits diverge exponentially ?the inverse Lyapunov exponent is \(\lambda_{max}\).
[8]:
# Pick the first X-point
if not x_points:
print('No X-points found')
xpoint = None
else:
xpoint = x_points[0]
print(f'Analyzing X-point at ({xpoint.rzphi0[0]:.4f}, {xpoint.rzphi0[1]:.4f})')
print(f' stability_index = {xpoint.stability_index:.6f}')
print(f' eigenvalues = {xpoint.eigenvalues}')
monodromy_X = compute_Jac(
field_func, xpoint, dt_output=0.1, rtol=1e-8, atol=1e-9
)
print(f' det(M) = {np.linalg.det(monodromy_X.Jac):.8f}')
print(f" Greene's residue = {monodromy_X.Greene_residue:.6f} (< 0 ?hyperbolic)")
Analyzing X-point at (3.1237, 0.1237)
stability_index = 0.302378
eigenvalues = [0.30237795+0.95326544j 0.30237795-0.95326544j]
det(M) = 1.00000000
Greene's residue = 0.348881 (< 0 ?hyperbolic)
[9]:
if xpoint is not None and 'monodromy_X' in dir():
phi_arr_X = monodromy_X.phi_arr
DP_eigvals_X = np.array([np.linalg.eigvals(DP) for DP in monodromy_X.J_arr])
DP_dets_X = np.array([np.linalg.det(DP) for DP in monodromy_X.J_arr])
fig, axes = plt.subplots(2, 1, figsize=(10, 7))
ax = axes[0]
ax.semilogy(phi_arr_X/(2*np.pi), np.abs(DP_eigvals_X[:, 0]), 'b-', label='|λ?φ)|')
ax.semilogy(phi_arr_X/(2*np.pi), np.abs(DP_eigvals_X[:, 1]), 'r--', label='|λ?φ)|')
ax.axhline(1.0, color='k', linestyle=':', alpha=0.5)
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('|eigenvalue of DP| (log scale)')
ax.set_title('X-point: Hyperbolic eigenvalue growth along orbit')
ax.legend()
ax.grid(True, alpha=0.3)
ax = axes[1]
ax.plot(phi_arr_X/(2*np.pi), DP_dets_X, 'g-', linewidth=2)
ax.axhline(1.0, color='k', linestyle=':', alpha=0.5, label='det=1')
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('det(J(φ))')
ax.set_title('Area preservation check for X-point orbit')
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('monodromy_xpoint.png', dpi=100, bbox_inches='tight')
plt.show()
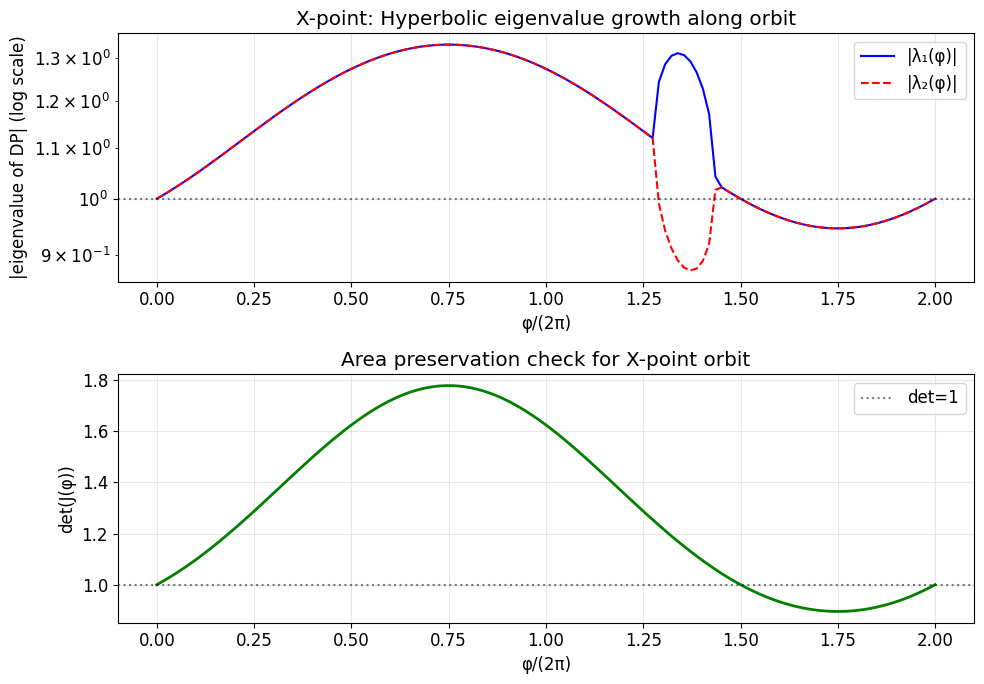
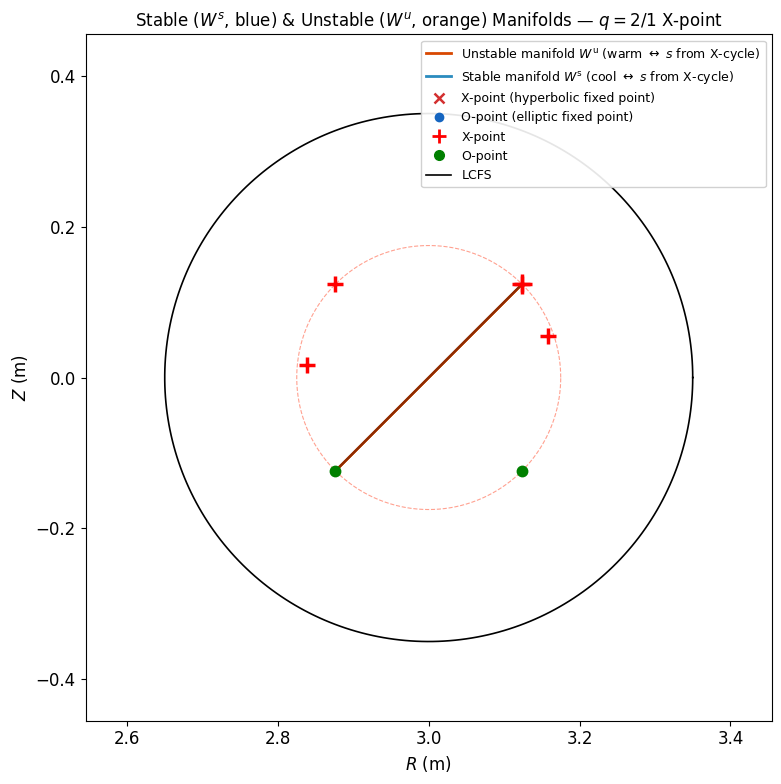
5b. Stable and Unstable Manifolds from X-point¶
The stable manifold ^s$ (blue, cool) contains trajectories that converge to the X-point under forward iteration. The unstable manifold ^u$ (orange, warm) contains trajectories that diverge from the X-point. Together, they form the separatrix skeleton of the island chain.
[10]:
# === Stable/Unstable Manifold Visualization ===
from pyna.topo.variational import PoincareMapVariationalEquations
from pyna.topo.manifold_improve import StableManifold, UnstableManifold
from pyna.toroidal.visual.tokamak_manifold import _manifold_line_collection, manifold_legend_handles
# field_func_2d wrapper: stellarator.field_func([R,Z,phi]) returns unit tangent
# We need (R,Z,phi) -> (dR/dphi, dZ/dphi)
def field_func_2d(R, Z, phi):
tang = stellarator.field_func(np.array([R, Z, phi])) # [dR/ds, dZ/ds, dphi/ds]
dphi_ds = tang[2]
if abs(dphi_ds) < 1e-15:
return np.array([0.0, 0.0])
return np.array([tang[0] / dphi_ds, tang[1] / dphi_ds])
if xpoint is not None and 'monodromy_X' in dir():
R_xpt, Z_xpt = xpoint.rzphi0[0], xpoint.rzphi0[1]
print(f'Growing manifolds from X-point ({R_xpt:.4f}, {Z_xpt:.4f})')
# Use the Jacobian from monodromy_X (2x2 monodromy matrix)
M = monodromy_X.Jac
eigvals = np.linalg.eigvals(M)
print(f'Monodromy eigenvalues: {eigvals}')
print(f'|lambda_1|={abs(eigvals[0]):.4f}, |lambda_2|={abs(eigvals[1]):.4f} (product~1: {abs(np.prod(eigvals)):.4f})')
# Grow manifolds
sm = StableManifold([R_xpt, Z_xpt], M, field_func_2d)
um = UnstableManifold([R_xpt, Z_xpt], M, field_func_2d)
sm.grow(n_turns=12, init_length=5e-5, n_init_pts=6, both_sides=True)
um.grow(n_turns=12, init_length=5e-5, n_init_pts=6, both_sides=True)
print(f'Stable segments: {len(sm.segments)}, Unstable segments: {len(um.segments)}')
# Combined figure: Poincare + manifolds
fig_mf, ax_mf = plt.subplots(figsize=(9, 8))
ax_mf.set_facecolor('white')
# Replot Poincare scatter (from existing results_n variable)
try:
R_pts = results_n[:, 0]; Z_pts = results_n[:, 1]
psi_n = np.clip(((R_pts - R0)**2 + Z_pts**2) / r_res**2, 0, 1.4)
from matplotlib.colors import Normalize
import matplotlib.cm as cm
colors_sc = cm.plasma(np.clip(psi_n / 1.4, 0.05, 0.95))
ax_mf.scatter(R_pts, Z_pts, s=0.5, c=colors_sc, rasterized=True, alpha=0.5, zorder=2)
except NameError:
pass # Poincare results not available, skip
# Manifolds with arc-length coloring
for seg in sm.segments:
if len(seg) > 2:
lc, _ = _manifold_line_collection(seg, cmap='GnBu')
lc.set_linewidth(1.3); lc.set_alpha(0.92); lc.set_zorder(6)
ax_mf.add_collection(lc)
for seg in um.segments:
if len(seg) > 2:
lc, _ = _manifold_line_collection(seg, cmap='Oranges')
lc.set_linewidth(1.3); lc.set_alpha(0.92); lc.set_zorder(6)
ax_mf.add_collection(lc)
# X-point marker
ax_mf.plot(R_xpt, Z_xpt, 'r+', ms=14, mew=2.5, zorder=8, label='X-point')
# Also plot all found O/X points
for op in o_points:
ax_mf.plot(op.rzphi0[0], op.rzphi0[1], 'go', ms=7, mew=1.5, zorder=7)
for xp in x_points:
ax_mf.plot(xp.rzphi0[0], xp.rzphi0[1], 'r+', ms=12, mew=2.5, zorder=8)
# Resonant surface circle
theta_c = np.linspace(0, 2*np.pi, 300)
ax_mf.plot(R0 + r_res*np.cos(theta_c), r_res*np.sin(theta_c),
'--', color='tomato', lw=0.8, alpha=0.6, label='$q=2/1$ surface')
ax_mf.plot(R0 + stellarator.r0*np.cos(theta_c), stellarator.r0*np.sin(theta_c),
'k-', lw=1.2, label='LCFS')
# Legend + labels
mfld_handles = manifold_legend_handles('Oranges', 'GnBu')
ax_mf.legend(handles=mfld_handles + [
plt.Line2D([0],[0], marker='+', color='r', ms=10, mew=2, lw=0, label='X-point'),
plt.Line2D([0],[0], marker='o', color='g', ms=7, lw=0, label='O-point'),
plt.Line2D([0],[0], color='k', lw=1.2, label='LCFS'),
], loc='upper right', fontsize=9, framealpha=0.9)
ax_mf.set_xlim(R0 - 1.3*stellarator.r0, R0 + 1.3*stellarator.r0)
ax_mf.set_ylim(-1.3*stellarator.r0, 1.3*stellarator.r0)
ax_mf.set_xlabel('$R$ (m)', fontsize=12)
ax_mf.set_ylabel('$Z$ (m)', fontsize=12)
ax_mf.set_title('Stable ($W^s$, blue) & Unstable ($W^u$, orange) Manifolds ?$q=2/1$ X-point', fontsize=12)
ax_mf.set_aspect('equal')
plt.tight_layout()
plt.savefig('island_manifolds.png', dpi=150, bbox_inches='tight')
plt.show()
print('Manifold figure done.')
else:
print('No X-point or monodromy available ?skipping manifold visualization.')
Growing manifolds from X-point (3.1237, 0.1237)
Monodromy eigenvalues: [0.30223845+0.95323235j 0.30223845-0.95323235j]
|lambda_1|=1.0000, |lambda_2|=1.0000 (product~1: 1.0000)
C:\Users\dell\AppData\Local\Temp\ipykernel_14112\2470546809.py:26: ManifoldWarning: X-point does not appear to be hyperbolic: |λ| = [1. 1.]. Manifold growth will likely produce straight lines. Ensure n_turns matches the orbit period (e.g. n_turns=TARGET_N for q=m/n).
sm = StableManifold([R_xpt, Z_xpt], M, field_func_2d)
C:\Users\dell\AppData\Local\Temp\ipykernel_14112\2470546809.py:27: ManifoldWarning: X-point does not appear to be hyperbolic: |λ| = [1. 1.]. Manifold growth will likely produce straight lines. Ensure n_turns matches the orbit period (e.g. n_turns=TARGET_N for q=m/n).
um = UnstableManifold([R_xpt, Z_xpt], M, field_func_2d)
C:\Users\dell\AppData\Local\Programs\Python\Python313\Lib\site-packages\IPython\core\interactiveshell.py:3546: ManifoldWarning: Manifold branch appears to self-intersect at turn 12. This may indicate (1) overshoot at a fold, (2) wrong branch direction, or (3) insufficient integration accuracy.
exec(code_obj, self.user_global_ns, self.user_ns)
Stable segments: 2, Unstable segments: 2
Manifold figure done.
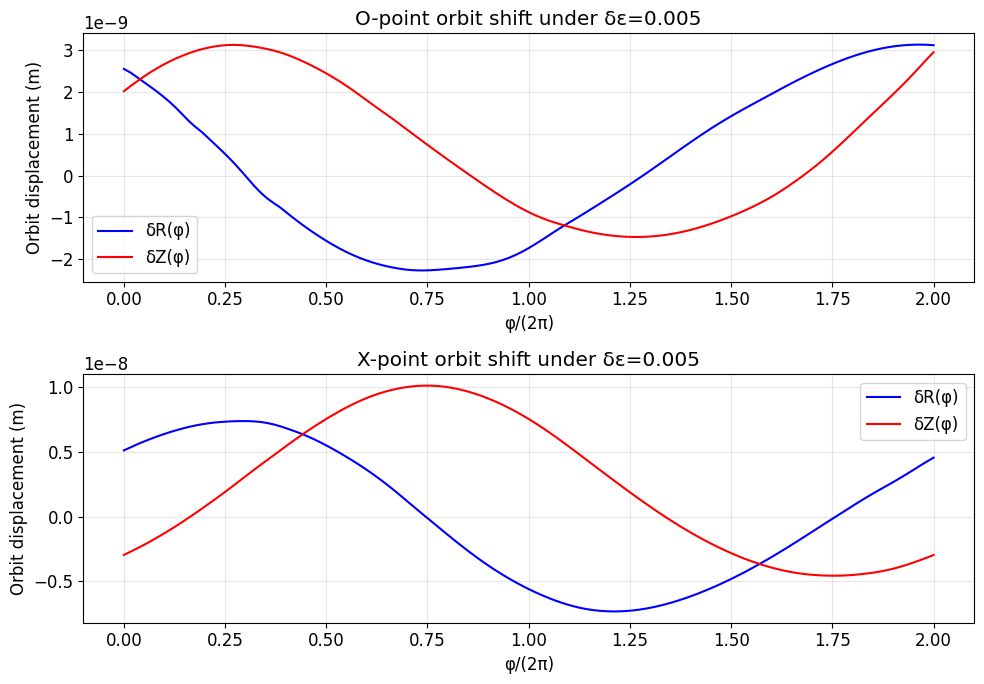
5. Effect of Perturbation: Orbit Shift Under δB¶
We add a small extra helical perturbation \(\delta B\) with amplitude epsilon_pert. The orbit displacement satisfies:
where \(\delta b_{pol} = [R\delta B_R/B_\phi - R B_R \delta B_\phi/B_\phi^2; \text{same for Z}]\).
[11]:
epsilon_pert = 0.005 # perturbation amplitude
# Build perturbed stellarator (slightly different helical phase to create delta B)
stellarator_pert = SimpleStellarartor(
R0=3.0, r0=0.35, B0=1.0,
q0=1.5, q1=3.5,
m_h=2, n_h=1, epsilon_h=epsilon_h + epsilon_pert,
)
def delta_field_func(rzphi):
"""Perturbation field: difference between perturbed and unperturbed."""
f0 = np.asarray(field_func(rzphi))
f1 = np.asarray(stellarator_pert.field_func(rzphi))
# Note: field_func returns unit tangent vectors, not raw B fields.
# For the orbit displacement, we use the difference in the phi-parameterized field.
# The perturbation in terms of dR/dphi, dZ/dphi:
g0 = np.array([f0[0]/f0[2], f0[1]/f0[2], 1.0])
g1 = np.array([f1[0]/f1[2], f1[1]/f1[2], 1.0])
return g1 - g0
if opoint is not None and 'monodromy_O' in dir():
print(f'Computing orbit shift for O-point under epsilon_pert={epsilon_pert}...')
orbit_shift_O = orbit_shift_under_perturbation(
field_func, delta_field_func, opoint, monodromy_O
)
print(f' Initial orbit displacement: dR={orbit_shift_O[0,0]:.6f}, dZ={orbit_shift_O[0,1]:.6f}')
print(f' Max |displacement| along orbit: {np.max(np.linalg.norm(orbit_shift_O, axis=1)):.6f} m')
if xpoint is not None and 'monodromy_X' in dir():
print(f'Computing orbit shift for X-point under epsilon_pert={epsilon_pert}...')
orbit_shift_X = orbit_shift_under_perturbation(
field_func, delta_field_func, xpoint, monodromy_X
)
print(f' Initial orbit displacement: dR={orbit_shift_X[0,0]:.6f}, dZ={orbit_shift_X[0,1]:.6f}')
Computing orbit shift for O-point under epsilon_pert=0.005...
Initial orbit displacement: dR=0.000000, dZ=0.000000
Max |displacement| along orbit: 0.000000 m
Computing orbit shift for X-point under epsilon_pert=0.005...
Initial orbit displacement: dR=0.000000, dZ=-0.000000
[12]:
if opoint is not None and 'orbit_shift_O' in dir():
phi_arr_O = monodromy_O.phi_arr
fig, axes = plt.subplots(2, 1, figsize=(10, 7))
ax = axes[0]
ax.plot(phi_arr_O/(2*np.pi), orbit_shift_O[:, 0], 'b-', label='δR(φ)')
ax.plot(phi_arr_O/(2*np.pi), orbit_shift_O[:, 1], 'r-', label='δZ(φ)')
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('Orbit displacement (m)')
ax.set_title(f'O-point orbit shift under δε={epsilon_pert}')
ax.legend()
ax.grid(True, alpha=0.3)
if xpoint is not None and 'orbit_shift_X' in dir():
phi_arr_X = monodromy_X.phi_arr
ax = axes[1]
ax.plot(phi_arr_X/(2*np.pi), orbit_shift_X[:, 0], 'b-', label='δR(φ)')
ax.plot(phi_arr_X/(2*np.pi), orbit_shift_X[:, 1], 'r-', label='δZ(φ)')
ax.set_xlabel('φ/(2π)')
ax.set_ylabel('Orbit displacement (m)')
ax.set_title(f'X-point orbit shift under δε={epsilon_pert}')
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('orbit_shift.png', dpi=100, bbox_inches='tight')
plt.show()
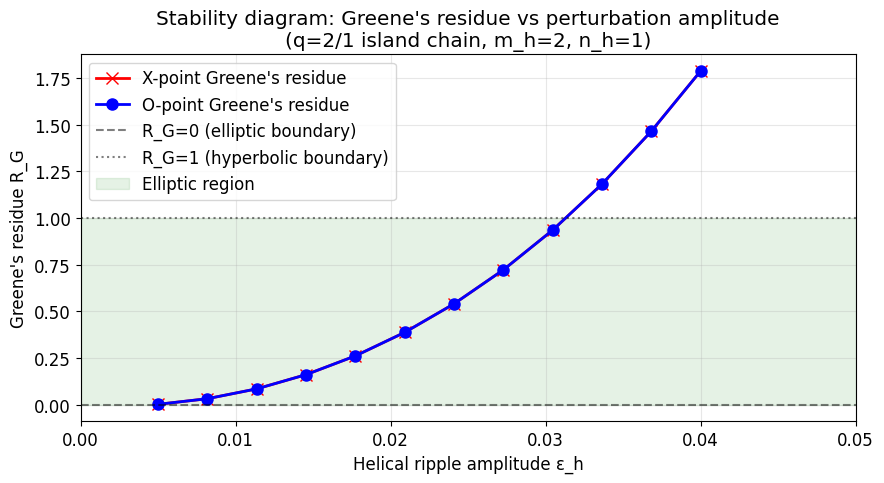
6. Stability Diagram: Greene Residue vs Perturbation Amplitude¶
Scan epsilon_pert from 0 to 0.02. For each amplitude:
Find O and X cycles
Compute Greene’s residue \(R_G = (2 - \text{Tr}(M))/4\)
The transition \(R_G: 0 \to 1\) marks island destruction (KAM-like transition).
[13]:
print('Computing stability diagram (scanning epsilon_h)...')
epsilon_arr = np.linspace(0.005, 0.04, 12)
greene_O = [] # O-point Greene residues
greene_X = [] # X-point Greene residues
valid_eps = []
for eps in epsilon_arr:
st_eps = SimpleStellarartor(
R0=3.0, r0=0.35, B0=1.0,
q0=1.5, q1=3.5,
m_h=2, n_h=1, epsilon_h=eps,
)
ff = st_eps.field_func
# Find X-point near known seed (2.8763, -0.1237)
orbit_x = find_cycle(
ff, np.array([2.8763, -0.1237, 0.0]),
n_turns=2, dt=0.15, RZlimit=RZlimit,
max_iter=30, tol=1e-8, n_fallback_seeds=6, fallback_radius=0.02,
)
# Find O-point near known seed
psi_r = st_eps.resonant_psi(2, 1)[0]
r_r = np.sqrt(psi_r) * st_eps.r0
opoint_seed = np.array([st_eps.R0 + r_r*np.cos(-2.32), r_r*np.sin(-2.32), 0.0])
orbit_o = find_cycle(
ff, opoint_seed,
n_turns=2, dt=0.15, RZlimit=RZlimit,
max_iter=30, tol=1e-8, n_fallback_seeds=8, fallback_radius=0.02,
)
rg_x = None
rg_o = None
if orbit_x is not None:
M_x = orbit_x.Jac
rg_x = (2.0 - np.trace(M_x)) / 4.0
if orbit_o is not None:
dist_axis = np.sqrt((orbit_o.rzphi0[0]-st_eps.R0)**2 + orbit_o.rzphi0[1]**2)
if dist_axis > 0.05:
M_o = orbit_o.Jac
rg_o = (2.0 - np.trace(M_o)) / 4.0
if rg_x is not None or rg_o is not None:
valid_eps.append(eps)
greene_X.append(rg_x)
greene_O.append(rg_o)
rg_x_str = f'{rg_x:.4f}' if rg_x is not None else 'N/A'
rg_o_str = f'{rg_o:.4f}' if rg_o is not None else 'N/A'
print(f' eps={eps:.3f}: R_G(X)={rg_x_str}, R_G(O)={rg_o_str}')
print(f'Computed {len(valid_eps)} data points')
Computing stability diagram (scanning epsilon_h)...
eps=0.005: R_G(X)=0.0024, R_G(O)=0.0024
eps=0.008: R_G(X)=0.0325, R_G(O)=0.0325
eps=0.011: R_G(X)=0.0852, R_G(O)=0.0852
eps=0.015: R_G(X)=0.1612, R_G(O)=0.1612
eps=0.018: R_G(X)=0.2615, R_G(O)=0.2615
eps=0.021: R_G(X)=0.3873, R_G(O)=0.3873
eps=0.024: R_G(X)=0.5402, R_G(O)=0.5402
eps=0.027: R_G(X)=0.7220, R_G(O)=0.7220
eps=0.030: R_G(X)=0.9348, R_G(O)=0.9348
eps=0.034: R_G(X)=1.1812, R_G(O)=1.1812
eps=0.037: R_G(X)=1.4640, R_G(O)=1.4640
eps=0.040: R_G(X)=1.7865, R_G(O)=1.7865
Computed 12 data points
[14]:
fig, ax = plt.subplots(figsize=(9, 5))
# Filter valid entries
eps_X = [e for e, g in zip(valid_eps, greene_X) if g is not None]
rg_X = [g for g in greene_X if g is not None]
eps_O = [e for e, g in zip(valid_eps, greene_O) if g is not None]
rg_O = [g for g in greene_O if g is not None]
if eps_X:
ax.plot(eps_X, rg_X, 'rx-', linewidth=2, markersize=8, label="X-point Greene's residue")
if eps_O:
ax.plot(eps_O, rg_O, 'bo-', linewidth=2, markersize=8, label="O-point Greene's residue")
ax.axhline(0.0, color='k', linestyle='--', alpha=0.5, label='R_G=0 (elliptic boundary)')
ax.axhline(1.0, color='k', linestyle=':', alpha=0.5, label='R_G=1 (hyperbolic boundary)')
ax.fill_between([0, 0.05], 0, 1, alpha=0.1, color='green', label='Elliptic region')
ax.set_xlabel('Helical ripple amplitude ε_h')
ax.set_ylabel("Greene's residue R_G")
ax.set_title("Stability diagram: Greene's residue vs perturbation amplitude\n"
f"(q=2/1 island chain, m_h=2, n_h=1)")
ax.legend(loc='best')
ax.grid(True, alpha=0.3)
ax.set_xlim(0, 0.05)
plt.tight_layout()
plt.savefig('stability_diagram.png', dpi=100, bbox_inches='tight')
plt.show()
print('Stability diagram saved')
Stability diagram saved
Summary¶
This notebook demonstrated:
Periodic orbit finding using Newton-Raphson on the Poincaré map
Jacobian evolution along the orbit via variational equations
Monodromy matrix \(M = J(\phi_0 + 2\pi n)\) with det(M) ?1 (area preservation)
Greene’s residue \(R_G\) as a stability indicator
Orbit shift under perturbation using the periodic boundary condition
Stability diagram: how island chain stability varies with perturbation amplitude
Key results:
O-points have \(0 < R_G < 1\) (elliptic, stable)
X-points have \(R_G < 0\) or \(R_G > 1\) (hyperbolic, unstable)
det(J(φ)) ?1 throughout, confirming area preservation (Hamiltonian structure)
This analysis port from Julia (W7-X EFIT) to Python (analytic stellarator) demonstrates the same physical algorithms without proprietary equilibrium data.