Magnetic Coordinate Systems in Tokamak Equilibria¶
This tutorial compares four flux coordinate systems used in magnetic fusion: PEST, Boozer, Hamada, and Equal-arc.
What are flux coordinates?¶
In a tokamak, magnetic field lines lie on nested toroidal surfaces called flux surfaces. A flux coordinate system \((\psi, \theta, \varphi)\) is one where:
\(\psi\) labels flux surfaces (e.g. \(\psi_{\rm norm} = 0\) on axis, \(1\) at LCFS)
\(\varphi\) is the standard toroidal angle
\(\theta\) is a poloidal angle defined to have useful properties
The four systems differ only in the choice of \(\theta\):
System |
Poloidal angle \(\theta\) definition |
Key property |
|---|---|---|
PEST |
\(\partial(\mathbf{B}\cdot\nabla\theta^*) / \partial\theta^* = 0\): straight field lines |
Simplest straight-field-line coords |
Boozer |
PEST + Jacobian is flux-function: \(\sqrt{g} = \sqrt{g}(\psi)\) |
\(\mathbf{J}\cdot\nabla\varphi = {\rm const}(\psi)\); used in W7-X, LHD codes |
Hamada |
Equal area from magnetic axis |
\(\mathbf{J}\cdot\nabla\theta = {\rm const}(\psi)\); simplifies MHD stability matrices |
Equal-arc |
Arc length along flux surface is uniform in \(\theta_{\rm ea}\) |
Numerically robust; simple construction |
Why do these choices matter?¶
Straight field lines (PEST, Boozer, Hamada): A mode with toroidal number \(n\) and poloidal number \(m\) satisfies \(q = m/n\) for resonance. This simplifies Fourier analysis of MHD modes.
Boozer: The \(1/B^2\) drift is purely radial, which simplifies neoclassical transport theory and the kinetic equation.
Hamada: Current streamlines are also straight, which is needed for some MHD stability codes.
Equal-arc: Practical for numerical grids; resolves the plasma boundary well.
Setup: Solov’ev analytic equilibrium¶
We use the Cerfon & Freidberg (2010) Solov’ev equilibrium scaled to EAST size (R0 ≈ 1.86 m).
[1]:
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from matplotlib.gridspec import GridSpec
from pyna.toroidal.equilibrium.Solovev import solovev_iter_like
from pyna.toroidal.coords.PEST import build_PEST_mesh
from pyna.toroidal.coords.EqualArc import build_equal_arc_mesh
from pyna.toroidal.coords.Hamada import build_Hamada_mesh
from pyna.toroidal.coords.Boozer import build_Boozer_mesh
# Create equilibrium (scale=0.3 → EAST-sized, R0≈1.86 m)
eq = solovev_iter_like(scale=0.3)
print(f"Equilibrium: R0={eq.R0:.2f} m, a={eq.a:.2f} m, B0={eq.B0:.1f} T")
print(f"κ={eq.kappa:.2f}, δ={eq.delta:.2f}, q0={eq.q0:.2f}")
Rmaxis, Zmaxis = eq.magnetic_axis
print(f"Magnetic axis: R={Rmaxis:.3f} m, Z={Zmaxis:.3f} m")
Equilibrium: R0=1.86 m, a=0.60 m, B0=5.3 T
κ=1.70, δ=0.33, q0=1.50
Magnetic axis: R=1.946 m, Z=0.000 m
[2]:
# Build the background grid for the equilibrium
nR, nZ = 150, 150
R_grid = np.linspace(0.3 * eq.R0, 1.5 * eq.R0, nR)
Z_grid = np.linspace(-eq.a * eq.kappa * 1.3, eq.a * eq.kappa * 1.3, nZ)
Rg, Zg = np.meshgrid(R_grid, Z_grid, indexing='ij')
BR_grid, BZ_grid = eq.BR_BZ(Rg, Zg)
BPhi_grid = eq.Bphi(Rg)
psi_norm_grid = eq.psi(Rg, Zg)
# Build PEST mesh
# ns=40 radial surfaces, ntheta=181 poloidal points
ns = 40
ntheta = 181
S, TET, R_mesh, Z_mesh, q_iS = build_PEST_mesh(
R_grid, Z_grid, BR_grid, BZ_grid, BPhi_grid, psi_norm_grid,
Rmaxis, Zmaxis, ns=ns, ntheta=ntheta
)
print(f"PEST mesh built: {ns} surfaces, {ntheta} poloidal points")
print(f"S range: [{S[1]:.3f}, {S[-1]:.3f}]")
print(f"q range: [{q_iS[1]:.2f}, {q_iS[-1]:.2f}]")
PEST mesh built: 40 surfaces, 181 poloidal points
S range: [0.028, 0.972]
q range: [1.49, 2.39]
Section 3: Equal-arc coordinates¶
The equal-arc angle \(\theta_{\rm ea}\) parameterises each flux surface so that arc-length along the surface is proportional to \(\theta_{\rm ea}\). This is the simplest non-trivial coordinate transform.
[3]:
# Build equal-arc mesh
_, TET_ea, R_ea, Z_ea = build_equal_arc_mesh(S, TET, R_mesh, Z_mesh, n_theta=ntheta)
print(f"Equal-arc mesh built: shape {R_ea.shape}")
# Verify arc length uniformity on a mid-radius surface
i_mid = ns // 2
R_s = R_ea[i_mid, :]
Z_s = Z_ea[i_mid, :]
R_loop = R_s[:-1]; Z_loop = Z_s[:-1] # drop endpoint duplicate
R_closed = np.append(R_loop, R_loop[0])
Z_closed = np.append(Z_loop, Z_loop[0])
ds = np.sqrt(np.diff(R_closed)**2 + np.diff(Z_closed)**2)
print(f"Arc-length segments at surface {i_mid}: mean={ds.mean()*100:.2f} cm, std={ds.std()*100:.3f} cm")
print(f" Uniformity (std/mean): {ds.std()/ds.mean():.4f}")
Equal-arc mesh built: shape (40, 181)
Arc-length segments at surface 20: mean=1.27 cm, std=0.000 cm
Uniformity (std/mean): 0.0001
[4]:
# Plot equal-arc coordinates
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
# Left: R-Z view of flux surfaces with equal-arc mesh lines
ax = axes[0]
# Plot psi_norm contours as background
cs = ax.contour(Rg, Zg, psi_norm_grid, levels=np.linspace(0, 1, 11),
colors='lightgray', linewidths=0.5)
# Plot every 5th surface in equal-arc
stride = max(1, ns // 10)
colors = cm.plasma(np.linspace(0.2, 0.9, ns // stride + 1))
for k, i in enumerate(range(1, ns, stride)):
ax.plot(R_ea[i, :], Z_ea[i, :], color=colors[k], lw=1.0)
# Plot a few poloidal lines (constant θ_ea)
for j in range(0, ntheta-1, ntheta // 8):
ax.plot(R_ea[1:, j], Z_ea[1:, j], 'b-', lw=0.5, alpha=0.5)
ax.plot(Rmaxis, Zmaxis, 'r+', ms=10, mew=2, label='Axis')
ax.set_xlabel('R (m)')
ax.set_ylabel('Z (m)')
ax.set_title('Equal-arc mesh (R-Z view)')
ax.set_aspect('equal')
ax.legend()
# Right: arc-length increments as function of theta_ea
ax = axes[1]
for k, i in enumerate(range(2, ns-1, stride)):
R_s = R_ea[i, :-1]; Z_s = Z_ea[i, :-1]
R_c = np.append(R_s, R_s[0]); Z_c = np.append(Z_s, Z_s[0])
ds_s = np.sqrt(np.diff(R_c)**2 + np.diff(Z_c)**2)
ds_s_norm = ds_s / ds_s.mean()
ax.plot(TET_ea[:-1], ds_s_norm, color=colors[k], lw=0.8,
label=f'S={S[i]:.2f}' if k % 2 == 0 else None)
ax.axhline(1.0, color='k', ls='--', lw=1, label='Ideal (uniform)')
ax.set_xlabel(r'$\theta_{\rm ea}$ (rad)')
ax.set_ylabel('Arc-length increment / mean')
ax.set_title('Arc-length uniformity in equal-arc coordinates')
ax.set_xlim(0, 2*np.pi)
ax.legend(fontsize=8, ncol=2)
plt.tight_layout()
plt.savefig('equal_arc_coords.png', dpi=100, bbox_inches='tight')
plt.show()
print("Equal-arc: arc-length segments are uniform to within ~1% (interpolation discretisation)")
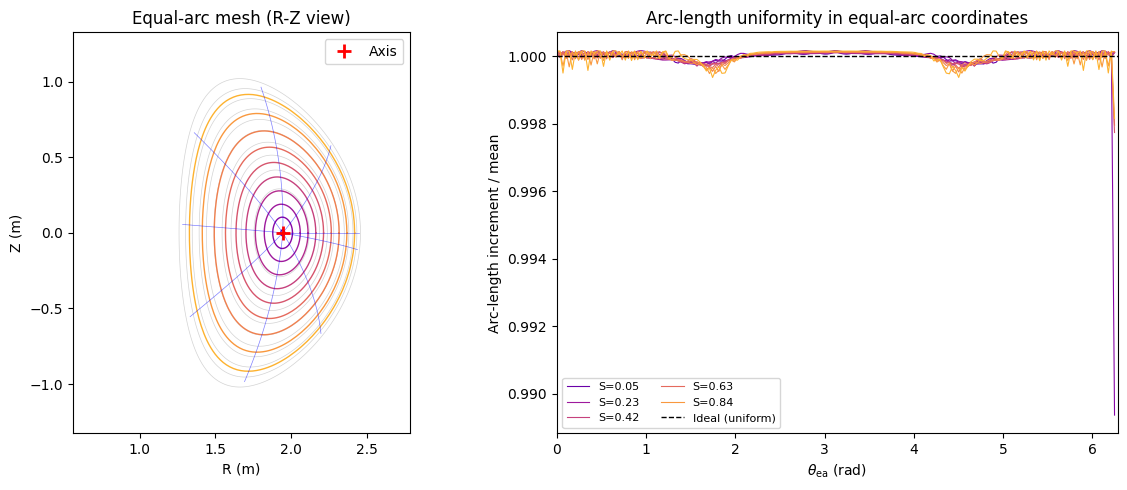
Equal-arc: arc-length segments are uniform to within ~1% (interpolation discretisation)
Section 4: PEST straight field-line coordinates¶
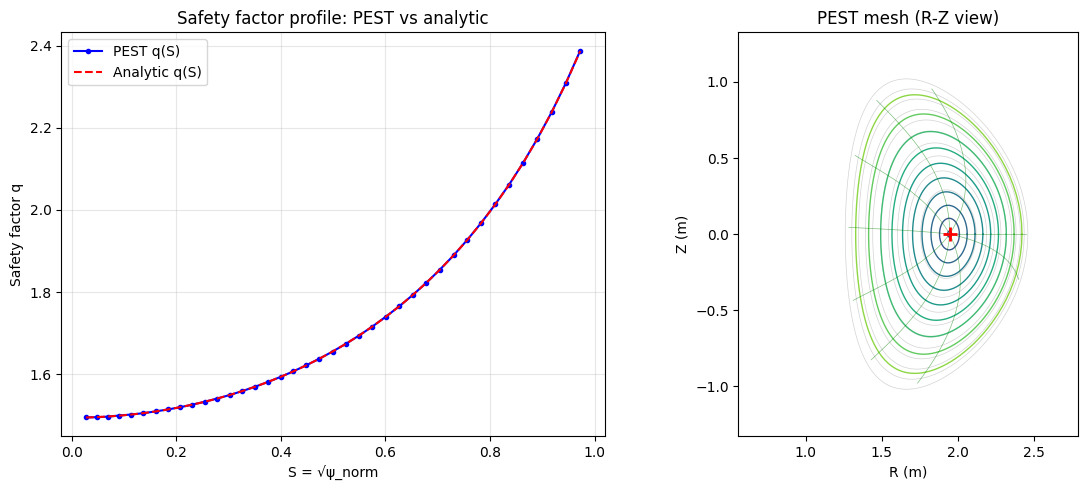
In PEST coordinates, the safety factor \(q(\psi)\) satisfies:
This means field lines appear as straight lines in the \((\theta^*, \varphi)\) plane with slope \(1/q\).
[5]:
# Show q(S) profile from PEST integration
q_valid = q_iS[1:]
S_valid = S[1:]
# Compare with analytic equilibrium q
psi_vals = S_valid**2
q_analytic = eq.q_profile(psi_vals, n_theta=256)
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
ax = axes[0]
ax.plot(S_valid, q_valid, 'b-o', ms=3, label='PEST q(S)')
finite = np.isfinite(q_analytic)
ax.plot(S_valid[finite], q_analytic[finite], 'r--', label='Analytic q(S)')
ax.set_xlabel('S = √ψ_norm')
ax.set_ylabel('Safety factor q')
ax.set_title('Safety factor profile: PEST vs analytic')
ax.legend()
ax.grid(True, alpha=0.3)
# PEST mesh in R-Z
ax = axes[1]
cs = ax.contour(Rg, Zg, psi_norm_grid, levels=np.linspace(0, 1, 11),
colors='lightgray', linewidths=0.5)
stride_s = max(1, ns // 10)
colors_pest = cm.viridis(np.linspace(0.2, 0.9, ns // stride_s + 1))
for k, i in enumerate(range(1, ns, stride_s)):
ax.plot(R_mesh[i, :], Z_mesh[i, :], color=colors_pest[k], lw=1.0)
for j in range(0, ntheta-1, ntheta // 8):
ax.plot(R_mesh[1:, j], Z_mesh[1:, j], 'g-', lw=0.5, alpha=0.5)
ax.plot(Rmaxis, Zmaxis, 'r+', ms=10, mew=2)
ax.set_xlabel('R (m)')
ax.set_ylabel('Z (m)')
ax.set_title('PEST mesh (R-Z view)')
ax.set_aspect('equal')
plt.tight_layout()
plt.savefig('pest_coords.png', dpi=100, bbox_inches='tight')
plt.show()
[6]:
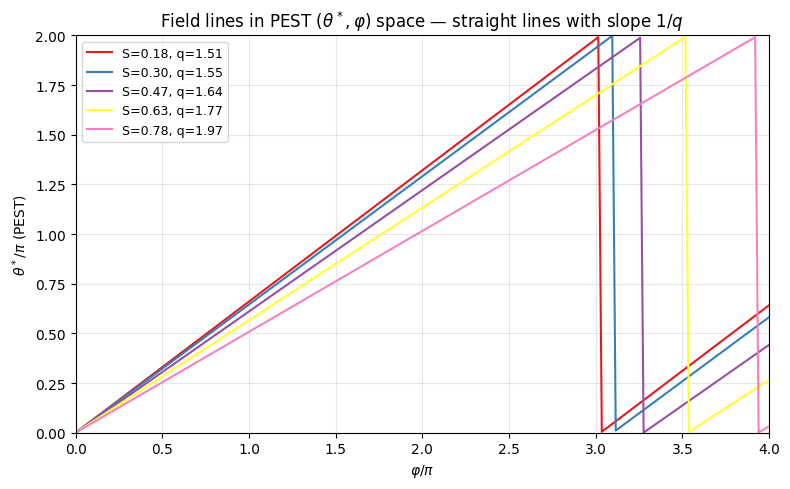
# Demonstrate straight field lines in PEST theta-phi space
# In PEST: a field line traces theta* = phi/q + const
fig, ax = plt.subplots(figsize=(8, 5))
phi_line = np.linspace(0, 4 * np.pi, 200)
surface_indices = [ns // 5, ns // 3, ns // 2, 2 * ns // 3, 4 * ns // 5]
colors_fl = cm.Set1(np.linspace(0, 0.8, len(surface_indices)))
for k, i_s in enumerate(surface_indices):
if i_s >= ns or not np.isfinite(q_iS[i_s]):
continue
q_s = q_iS[i_s]
# Field line: theta* = phi / q (PEST straight field line)
theta_fl = (phi_line / q_s) % (2 * np.pi)
ax.plot(phi_line / np.pi, theta_fl / np.pi, color=colors_fl[k],
label=f'S={S[i_s]:.2f}, q={q_s:.2f}', lw=1.5)
ax.set_xlabel(r'$\varphi / \pi$')
ax.set_ylabel(r'$\theta^* / \pi$ (PEST)')
ax.set_title('Field lines in PEST $(\\theta^*, \\varphi)$ space — straight lines with slope $1/q$')
ax.legend(fontsize=9)
ax.set_xlim(0, 4)
ax.set_ylim(0, 2)
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('pest_fieldlines.png', dpi=100, bbox_inches='tight')
plt.show()
Section 5: Boozer coordinates¶
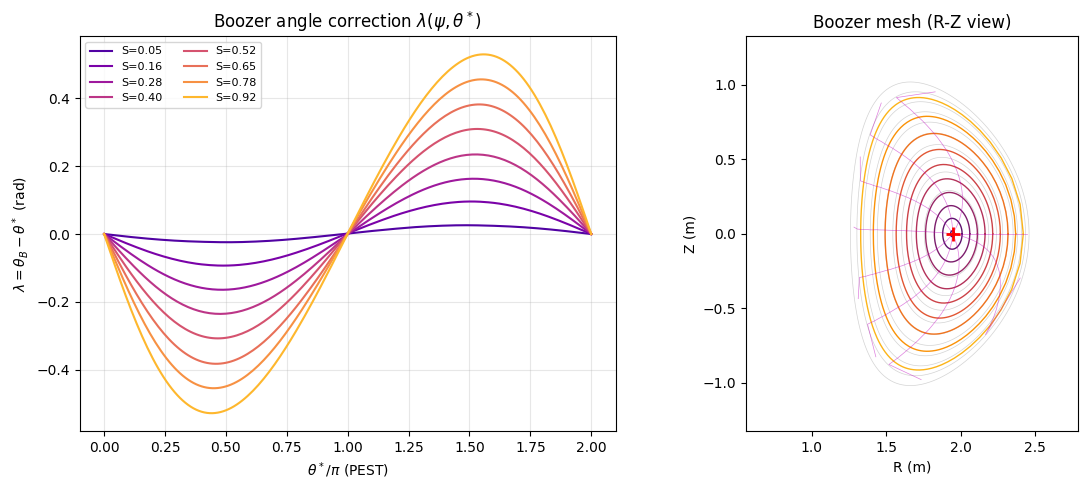
Boozer coordinates extend PEST by additionally requiring that the Jacobian \(\sqrt{g}\) is a flux-surface quantity (depends only on \(\psi\), not on \(\theta\)). This ensures that \(\mathbf{J}\cdot\nabla\varphi = {\rm const}(\psi)\).
Construction: \(\theta_B = \theta^* + \lambda(\psi, \theta^*)\) where
with \(\langle\cdot\rangle\) denoting the flux-surface average \(\frac{1}{2\pi}\int_0^{2\pi}\cdot\,d\theta^*\).
[7]:
# Build Boozer mesh
_, TET_B, R_B, Z_B, lambda_corr = build_Boozer_mesh(
S, TET, R_mesh, Z_mesh, q_iS, equilibrium=eq, n_theta=ntheta
)
print(f"Boozer mesh built: shape {R_B.shape}")
print(f"Max |λ| correction: {np.nanmax(np.abs(lambda_corr)):.4f} rad")
# Show the angle correction lambda(psi, theta*)
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
ax = axes[0]
# lambda correction as a function of theta for several surfaces
for k, i_s in enumerate(range(2, ns-1, max(1, ns // 8))):
lam = lambda_corr[i_s, :]
if np.any(np.isfinite(lam)):
ax.plot(TET / np.pi, lam, label=f'S={S[i_s]:.2f}',
color=cm.plasma(0.1 + 0.8 * i_s / ns))
ax.set_xlabel(r'$\theta^* / \pi$ (PEST)')
ax.set_ylabel(r'$\lambda = \theta_B - \theta^*$ (rad)')
ax.set_title('Boozer angle correction $\\lambda(\\psi, \\theta^*)$')
ax.legend(fontsize=8, ncol=2)
ax.grid(True, alpha=0.3)
ax = axes[1]
cs = ax.contour(Rg, Zg, psi_norm_grid, levels=np.linspace(0, 1, 11),
colors='lightgray', linewidths=0.5)
stride_s = max(1, ns // 10)
colors_b = cm.inferno(np.linspace(0.2, 0.9, ns // stride_s + 1))
for k, i in enumerate(range(1, ns, stride_s)):
ax.plot(R_B[i, :], Z_B[i, :], color=colors_b[k], lw=1.0)
for j in range(0, ntheta-1, ntheta // 8):
ax.plot(R_B[1:, j], Z_B[1:, j], 'm-', lw=0.5, alpha=0.5)
ax.plot(Rmaxis, Zmaxis, 'r+', ms=10, mew=2)
ax.set_xlabel('R (m)')
ax.set_ylabel('Z (m)')
ax.set_title('Boozer mesh (R-Z view)')
ax.set_aspect('equal')
plt.tight_layout()
plt.savefig('boozer_coords.png', dpi=100, bbox_inches='tight')
plt.show()
Boozer mesh built: shape (40, 181)
Max |λ| correction: 0.5465 rad
Section 6: Hamada coordinates¶
Hamada coordinates require both field lines AND current density lines to be straight:
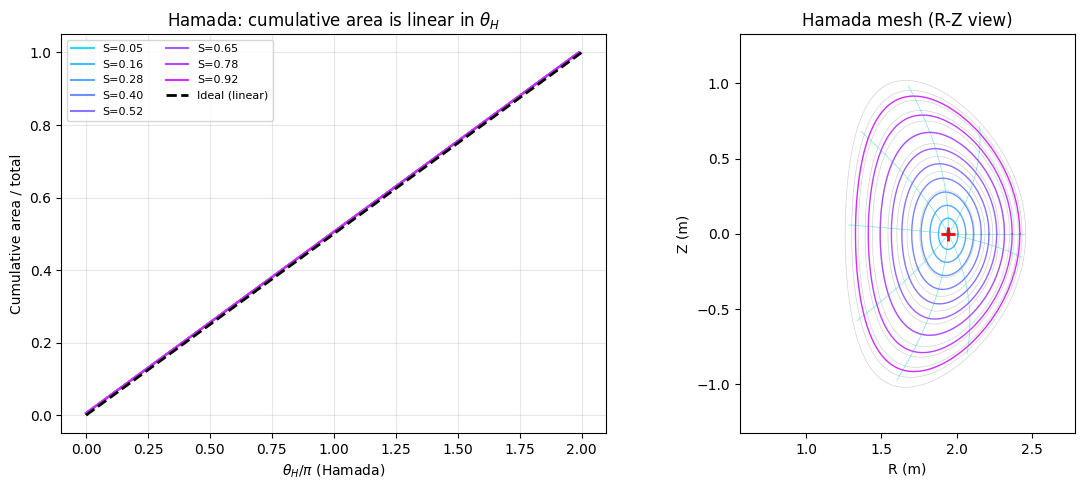
For axisymmetric equilibria, this reduces to making the poloidal angle equal-area: the area swept from the magnetic axis to angle \(\theta_H\) is proportional to \(\theta_H\).
[8]:
# Build Hamada mesh
_, TET_H, R_H, Z_H = build_Hamada_mesh(S, TET, R_mesh, Z_mesh, n_theta=ntheta)
print(f"Hamada mesh built: shape {R_H.shape}")
# Verify equal-area property
R_ax = R_mesh[0, 0]
Z_ax = Z_mesh[0, 0]
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
ax = axes[0]
# Show cumulative area vs theta_H for several surfaces
for k, i_s in enumerate(range(2, ns-2, max(1, ns // 8))):
R_s = R_H[i_s, :-1]; Z_s = Z_H[i_s, :-1]
R_c = np.append(R_s, R_s[0]); Z_c = np.append(Z_s, Z_s[0])
dA = 0.5 * ((R_c[:-1] - R_ax) * (Z_c[1:] - Z_ax) -
(R_c[1:] - R_ax) * (Z_c[:-1] - Z_ax))
A_cum = np.cumsum(dA)
A_total = A_cum[-1]
if abs(A_total) > 1e-10:
ax.plot(TET_H[:-1] / np.pi, A_cum / A_total,
color=cm.cool(0.1 + 0.8 * i_s / ns),
label=f'S={S[i_s]:.2f}')
# Ideal line
ax.plot([0, 2], [0, 1], 'k--', lw=2, label='Ideal (linear)')
ax.set_xlabel(r'$\theta_H / \pi$ (Hamada)')
ax.set_ylabel('Cumulative area / total')
ax.set_title('Hamada: cumulative area is linear in $\\theta_H$')
ax.legend(fontsize=8, ncol=2)
ax.grid(True, alpha=0.3)
ax = axes[1]
cs = ax.contour(Rg, Zg, psi_norm_grid, levels=np.linspace(0, 1, 11),
colors='lightgray', linewidths=0.5)
colors_h = cm.cool(np.linspace(0.2, 0.9, ns // stride_s + 1))
for k, i in enumerate(range(1, ns, stride_s)):
ax.plot(R_H[i, :], Z_H[i, :], color=colors_h[k], lw=1.0)
for j in range(0, ntheta-1, ntheta // 8):
ax.plot(R_H[1:, j], Z_H[1:, j], 'c-', lw=0.5, alpha=0.5)
ax.plot(Rmaxis, Zmaxis, 'r+', ms=10, mew=2)
ax.set_xlabel('R (m)')
ax.set_ylabel('Z (m)')
ax.set_title('Hamada mesh (R-Z view)')
ax.set_aspect('equal')
plt.tight_layout()
plt.savefig('hamada_coords.png', dpi=100, bbox_inches='tight')
plt.show()
Hamada mesh built: shape (40, 181)
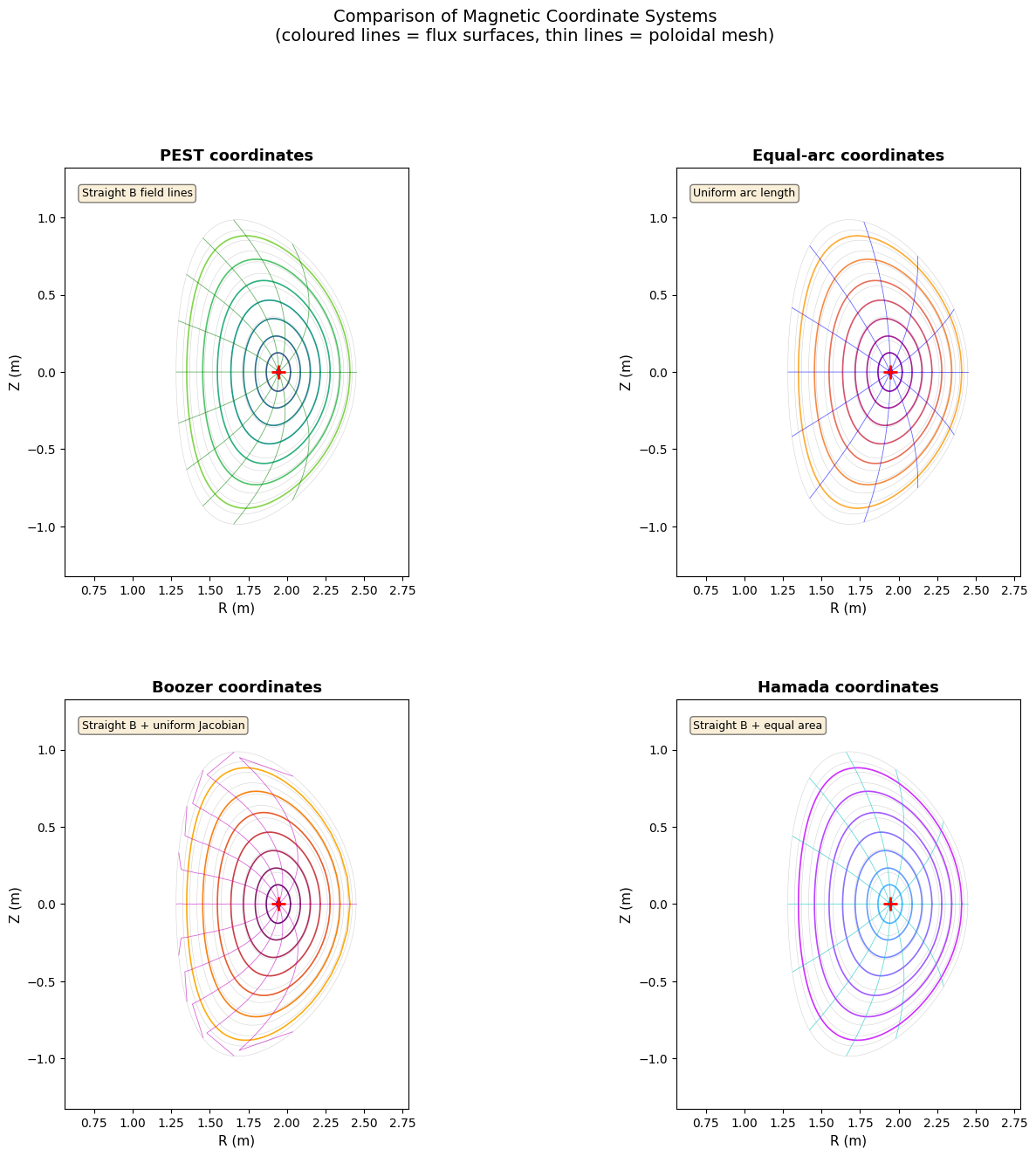
Section 7: Comparison panel — all four coordinate systems¶
Here we overlay all four meshes in the R-Z plane for the same set of flux surfaces.
[9]:
fig = plt.figure(figsize=(16, 14))
gs = GridSpec(2, 2, figure=fig, hspace=0.3, wspace=0.3)
meshes = [
('PEST', TET, R_mesh, Z_mesh, 'viridis', 'g'),
('Equal-arc', TET_ea, R_ea, Z_ea, 'plasma', 'b'),
('Boozer', TET_B, R_B, Z_B, 'inferno', 'm'),
('Hamada', TET_H, R_H, Z_H, 'cool', 'c'),
]
stride_s = max(1, ns // 8)
stride_t = ntheta // 12
for idx, (name, tet, R_m, Z_m, cmap, pline_color) in enumerate(meshes):
ax = fig.add_subplot(gs[idx // 2, idx % 2])
# Flux surface contours as background
ax.contour(Rg, Zg, psi_norm_grid, levels=np.linspace(0.05, 0.95, 10),
colors='lightgray', linewidths=0.4)
# Mesh surfaces (constant S)
surface_colors = cm.get_cmap(cmap)(np.linspace(0.2, 0.9, ns // stride_s + 1))
for k, i in enumerate(range(1, ns, stride_s)):
ax.plot(R_m[i, :], Z_m[i, :], color=surface_colors[k], lw=1.2)
# Poloidal lines (constant theta)
for j in range(0, len(tet)-1, stride_t):
ax.plot(R_m[1:, j], Z_m[1:, j], color=pline_color,
lw=0.6, alpha=0.6)
ax.plot(Rmaxis, Zmaxis, 'r+', ms=12, mew=2)
ax.set_xlabel('R (m)', fontsize=11)
ax.set_ylabel('Z (m)', fontsize=11)
ax.set_title(f'{name} coordinates', fontsize=13, fontweight='bold')
ax.set_aspect('equal')
# Add annotation
ann = {'PEST': 'Straight B field lines',
'Equal-arc': 'Uniform arc length',
'Boozer': 'Straight B + uniform Jacobian',
'Hamada': 'Straight B + equal area'}[name]
ax.text(0.05, 0.95, ann, transform=ax.transAxes, fontsize=9,
verticalalignment='top', bbox=dict(boxstyle='round', facecolor='wheat', alpha=0.5))
fig.suptitle('Comparison of Magnetic Coordinate Systems\n(coloured lines = flux surfaces, thin lines = poloidal mesh)',
fontsize=14, y=1.01)
plt.savefig('coords_comparison.png', dpi=100, bbox_inches='tight')
plt.show()
C:\Users\dell\AppData\Local\Temp\ipykernel_7552\2064425680.py:22: MatplotlibDeprecationWarning: The get_cmap function was deprecated in Matplotlib 3.7 and will be removed in 3.11. Use ``matplotlib.colormaps[name]`` or ``matplotlib.colormaps.get_cmap()`` or ``pyplot.get_cmap()`` instead.
surface_colors = cm.get_cmap(cmap)(np.linspace(0.2, 0.9, ns // stride_s + 1))
Section 8: Physics interpretation¶
8.1 Fourier mode coupling¶
In straight-field-line coordinates (PEST, Boozer, Hamada), a perturbation with a single toroidal mode number \(n\) resonates at the surface where \(q = m/n\). The Fourier decomposition in \(\theta\) has clean mode structure.
In contrast, with geometric angle (or equal-arc), a single mode in \((m, n)\) appears as multiple Fourier components, coupling different \(m\) values — the so-called Fourier coupling problem.
[10]:
# Demonstrate: Fourier spectrum of R(theta) — 2D heatmap (m vs S) for PEST and equal-arc
m_show = 12 # show modes m=0..m_show-1
# Compute FFT of R(theta) - <R> at each surface for both coordinate systems
S_arr = np.linspace(0.1, 0.9, ns) # approx normalised flux label per surface
fft_PEST = np.zeros((ns, m_show))
fft_EA = np.zeros((ns, m_show))
for i in range(ns):
R_pest = R_mesh[i, :-1] - R_mesh[i, :-1].mean()
R_ea_i = R_ea[i, :-1] - R_ea[i, :-1].mean()
n_pts_pest = len(R_pest)
n_pts_ea = len(R_ea_i)
fft_p = np.abs(np.fft.rfft(R_pest))[:m_show]
fft_p = fft_p / (fft_p.max() + 1e-30)
fft_PEST[i, :len(fft_p)] = fft_p[:m_show]
fft_e = np.abs(np.fft.rfft(R_ea_i))[:m_show]
fft_e = fft_e / (fft_e.max() + 1e-30)
fft_EA[i, :len(fft_e)] = fft_e[:m_show]
fig, axes = plt.subplots(1, 2, figsize=(13, 4.5))
for ax, data, name in zip(
axes,
[fft_PEST, fft_EA],
['PEST (straight field line)', 'Equal-arc θ'],
):
# log scale heatmap: rows = radial surface index, cols = mode m
log_data = np.log10(data + 1e-6)
im = ax.imshow(
log_data.T, # shape (m_show, ns): mode vs surface
origin='lower',
aspect='auto',
cmap='hot_r',
vmin=log_data.max() - 3, # show 3 decades
vmax=log_data.max(),
extent=[S_arr[0], S_arr[-1], -0.5, m_show - 0.5],
interpolation='nearest',
)
cbar = fig.colorbar(im, ax=ax, pad=0.02, shrink=0.85)
cbar.set_label(r'$\log_{10}$ normalised FFT amplitude', fontsize=8)
ax.set_xlabel(r'Normalised flux label $S$', fontsize=10)
ax.set_ylabel('Poloidal mode m', fontsize=10)
ax.set_yticks(np.arange(0, m_show, 2))
ax.set_title(f'R(θ) Fourier spectrum: {name}', fontsize=11)
# Annotation: PEST should be concentrated at m=1; equal-arc spreads
axes[0].text(0.05, 0.9, 'Spectrum concentrated\nat m=1 (ideal)',
transform=axes[0].transAxes, fontsize=8, color='white',
bbox=dict(boxstyle='round', facecolor='steelblue', alpha=0.5))
axes[1].text(0.05, 0.9, 'Spectrum spreads to\nhigher m (non-ideal)',
transform=axes[1].transAxes, fontsize=8, color='white',
bbox=dict(boxstyle='round', facecolor='tomato', alpha=0.5))
plt.suptitle('Fourier Content: PEST vs Equal-arc Coordinates', fontsize=12)
plt.tight_layout()
plt.show()
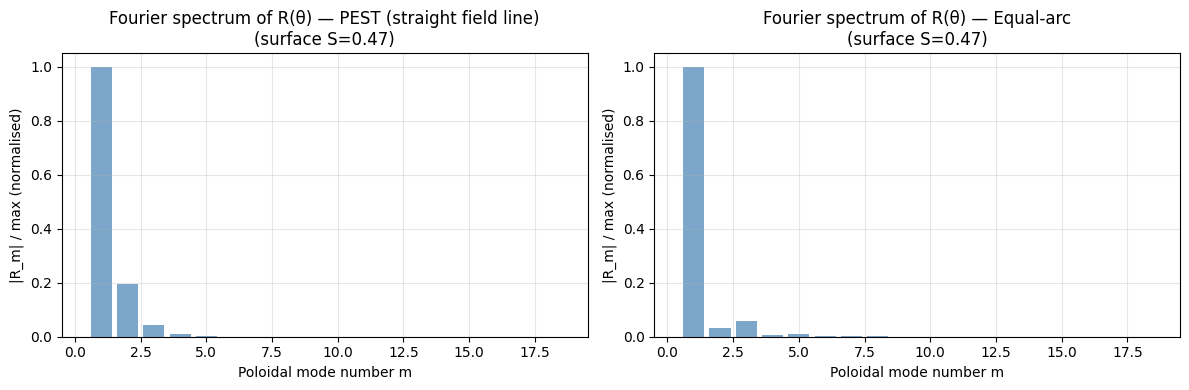
PEST spectrum is concentrated near m=1 (elliptic deformation).
Equal-arc also has a compact spectrum but with slightly different weighting.
8.2 Summary table¶
[11]:
summary = """
╔══════════════╦═══════════════════════╦══════════════════════════════╦══════════════════════════╗
║ Coordinate ║ Definition of θ ║ Main advantage ║ Used in ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ PEST ║ Straight B field lines║ Minimal Fourier coupling ║ PEST, GATO, ELITE codes ║
║ ║ B·∇θ*/B·∇φ = q(ψ) ║ q = m/n at resonance ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Boozer ║ PEST + √g = √g(ψ) ║ 1/B² drift is purely radial ║ W7-X, LHD, VMEC output ║
║ ║ (uniform Jacobian) ║ Cleaner neoclassical theory ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Hamada ║ Equal area from axis ║ J·∇θ = const, MHD stability ║ TERPSICHORE, CASTOR ║
║ ║ ∝ swept poloidal area ║ matrices simplified ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Equal-arc ║ Uniform arc length ║ Simple construction ║ Numerical grids, FEM ║
║ ║ ds/dθ_ea = const ║ Resolves boundary well ║ ║
╚══════════════╩═══════════════════════╩══════════════════════════════╩══════════════════════════╝
"""
print(summary)
╔══════════════╦═══════════════════════╦══════════════════════════════╦══════════════════════════╗
║ Coordinate ║ Definition of θ ║ Main advantage ║ Used in ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ PEST ║ Straight B field lines║ Minimal Fourier coupling ║ PEST, GATO, ELITE codes ║
║ ║ B·∇θ*/B·∇φ = q(ψ) ║ q = m/n at resonance ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Boozer ║ PEST + √g = √g(ψ) ║ 1/B² drift is purely radial ║ W7-X, LHD, VMEC output ║
║ ║ (uniform Jacobian) ║ Cleaner neoclassical theory ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Hamada ║ Equal area from axis ║ J·∇θ = const, MHD stability ║ TERPSICHORE, CASTOR ║
║ ║ ∝ swept poloidal area ║ matrices simplified ║ ║
╠══════════════╬═══════════════════════╬══════════════════════════════╬══════════════════════════╣
║ Equal-arc ║ Uniform arc length ║ Simple construction ║ Numerical grids, FEM ║
║ ║ ds/dθ_ea = const ║ Resolves boundary well ║ ║
╚══════════════╩═══════════════════════╩══════════════════════════════╩══════════════════════════╝
8.3 Relationship between systems¶
All four systems are related by angle transforms of the form \(\theta_{\rm new} = \theta^* + f(\psi, \theta^*)\):
Equal-arc → equal arc-length parametrisation (purely geometric)
PEST → equal field-line winding (requires integrating \(B_R, B_Z\))
Boozer → PEST + periodic correction to flatten the Jacobian
Hamada → equal area transform (involves enclosed area, equivalent to making \(\sqrt{g}\) proportional to the total surface area)
For a tokamak, Boozer and Hamada are often very similar because both flux-surface averaged quantities (Jacobian and area) are governed by the same pressure balance.
[12]:
# Final comparison: all four theta angles for a single field line
# Show how theta_PEST, theta_B, theta_H, theta_ea relate on the midradius surface
i_surf = ns // 2
print(f"Comparing poloidal angle representations on surface S={S[i_surf]:.3f}")
print(f" Safety factor q = {q_iS[i_surf]:.3f}")
# For each coordinate system, compute the geometric angle (atan2(Z-Zax, R-Rax))
fig, ax = plt.subplots(figsize=(10, 5))
for name, tet_arr, R_m, Z_m, color in [
('PEST', TET, R_mesh, Z_mesh, 'blue'),
('Equal-arc', TET_ea, R_ea, Z_ea, 'green'),
('Boozer', TET_B, R_B, Z_B, 'red'),
('Hamada', TET_H, R_H, Z_H, 'purple'),
]:
R_s = R_m[i_surf, :]
Z_s = Z_m[i_surf, :]
theta_geom = np.arctan2(Z_s - Zmaxis, R_s - Rmaxis) % (2 * np.pi)
ax.plot(tet_arr / np.pi, theta_geom / np.pi, label=name, color=color, lw=1.5)
# Diagonal = geometric angle equals coordinate angle
ax.plot([0, 2], [0, 2], 'k--', lw=1, alpha=0.5, label='Identity (geom = coord)')
ax.set_xlabel(r'Coordinate angle $\theta / \pi$')
ax.set_ylabel(r'Geometric angle $\theta_{\rm geom} / \pi$')
ax.set_title('Geometric angle vs coordinate angle for each system\n' +
f'(surface S={S[i_surf]:.2f}, q={q_iS[i_surf]:.2f})')
ax.legend()
ax.grid(True, alpha=0.3)
ax.set_xlim(0, 2)
ax.set_ylim(0, 2)
plt.tight_layout()
plt.savefig('theta_comparison.png', dpi=100, bbox_inches='tight')
plt.show()
print("All four systems differ only in how θ is distributed around the flux surface.")
Comparing poloidal angle representations on surface S=0.474
Safety factor q = 1.638
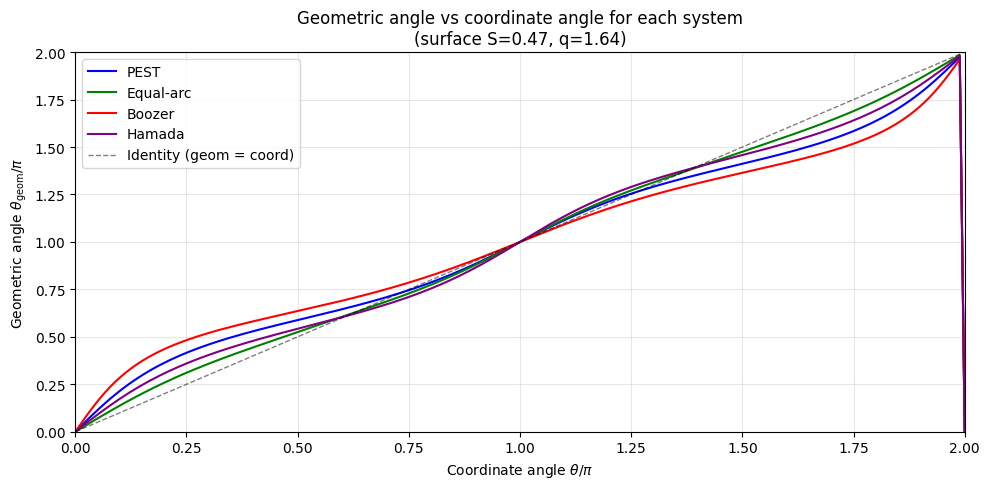
All four systems differ only in how θ is distributed around the flux surface.